Jump to:
Project Summary
Economic challenges imposed by climate change and disease on the aquaculture industry necessitate advances for improved animal welfare and resiliency. Biomarkers associated with environmental and disease resilience traits can be leveraged in breeding and management strategies. However, their discovery has been limited in part by the complexity of molecular systems and the cost of genomics tools used to understand them. Advances in computational approaches including machine learning algorithms, together with the wealth of genomic data that has amassed, enable powerful meta-analyses for improved biomarker discovery in aquaculture species.
This project aims to advance the discovery and characterization of biomarkers through mining publicly available shellfish genomic datasets from resilient populations.
The objectives are to:
1) Develop standardized open-access, user-friendly, reproducible bioinformatics pipelines for resilience biomarker discovery through systematic reanalysis, data integration and meta-analysis
2) Build a user-friendly open-access comprehensive database of candidate resilience biomarkers that is widely available for use by the aquaculture community.
The resulting database will enable improved molecular tool development for more efficient phenotype selection and health monitoring, implementation of selection methods that use a systems biology approach for simultaneous improvement of multiple traits, and ultimately increased animal fitness and improved animal welfare.
View more project details here
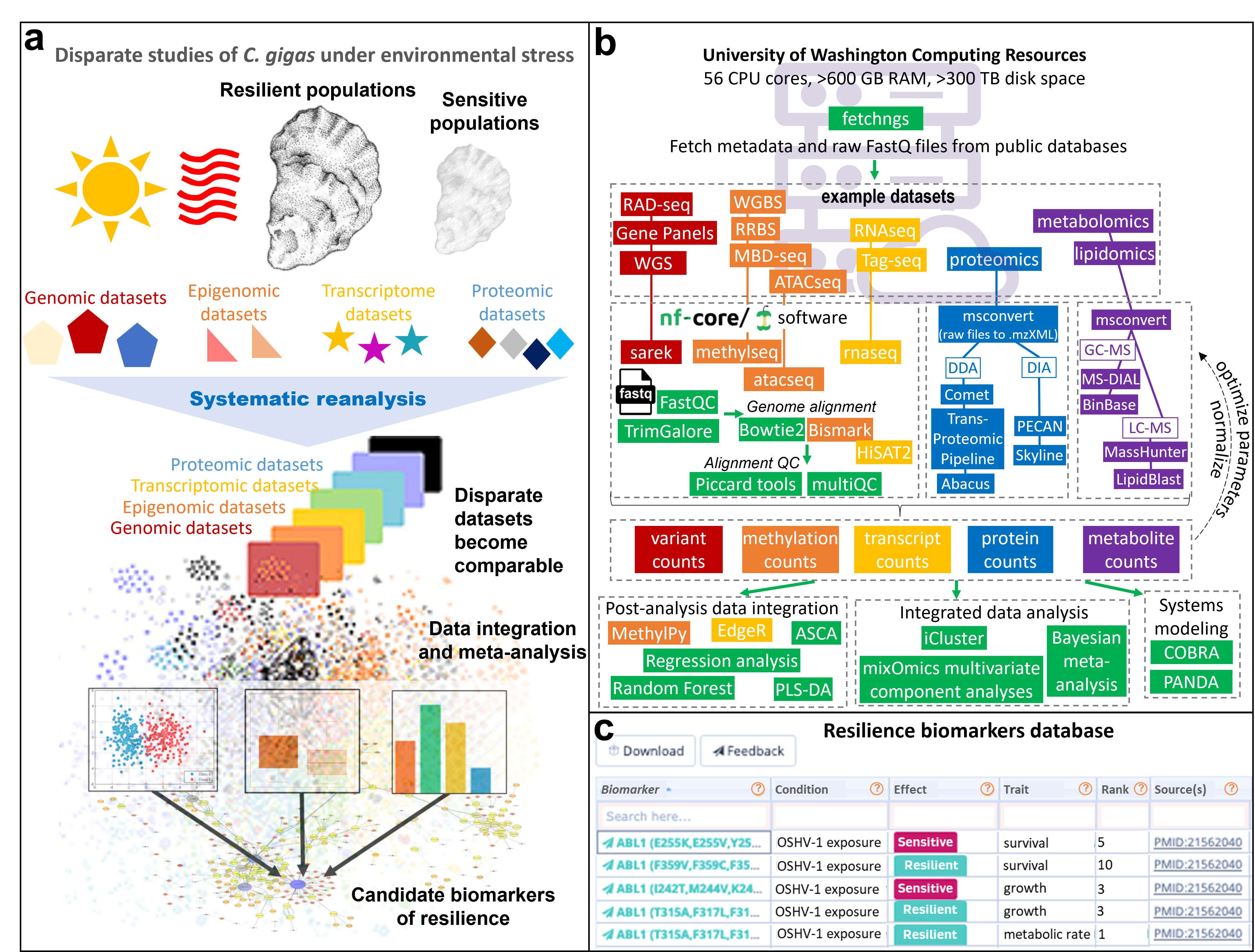 a) Graphical summary of Objective 1. b) Example pipelines and software for systematic reanalysis and data integration. c) Example of resilience biomarker database (modeled after Tamborero et al. 201846).
a) Graphical summary of Objective 1. b) Example pipelines and software for systematic reanalysis and data integration. c) Example of resilience biomarker database (modeled after Tamborero et al. 201846).
Omics datasets processed
Biomarker Database
People
Project Director:
Shelly Wanamaker, PhD (she/her)
Research Scientist
shelly.wanamaker@gmgi.org
https://github.com/shellywanamaker
Project Personnel:
Emma Strand, PhD (she/her)
Postdoctoral Scientist
emma.strand@gmgi.org
https://github.com/emmastrand
Steve Yost (he/him)
Bioinformatics Software Engineer
steve.yost+gmgi@gmail.com
Office and mailing address:
Gloucester Marine Genomics Institute
417 Main Street
Gloucester, MA 01930 USA
Funding:
This project is supported by the USDA Agriculture and Food Research Initiative Animal Health and Production and Animal Products program under the Animal Breeding, Genetics, and Genomics section award number 2024-67015-41794 to Dr. Shelly A. Wanamaker.